Representation Learning through Matrix Factorization
Principal Component Analysis (PCA) and Singular Value Decomposition (SVD) has been around for a while, and have been successfully utilized for learning intermediary representations of data for quite some time. This post will be recap on what they actually do, and how they work.
Note: this post is part 2 of my Representation Learning series. See part 1 and part 1.5.
Matrix Factorization
Matrix factorization is a way of decomposing any matrix $M$ into multiple matrices. Often this is done for two reasons: in order to reduce the matrix to a lower rank while keeping as much information as possible (dimensionality reduction), or to expose some underlying factors that appear naturally in the data. As this is a series about representation learning, we will be mostly concerned about exposing the underlying factors.
In the context of Machine Learning, it’s a good idea to think of the matrix $M$ as your data matrix, with $n$ data points and $m$ features. This can be the values you get from a database table or a csv file, or a tf-idf representation of documents, or anything that you have previously translated into a matrix of numbers.
It’s easy to ignore this, but I find it useful to acknowledge that if you had to translate your data into a matrix of numbers, you’ve already done some sort of Representation Learning already. So congrats! You’re now doing Hierarchical Representation Learning, which is a powerful tool that’s going to be covered more in-depth in the future posts.
This post will deal with Linear Algebra concepts such as eigenvectors and eigenvalues. If you need a refresher, I highly suggest checking the Essence of linear algebra series up in Youtube, which gives a very intuitive and graphical explanation of the concepts.
PCA
PCA is an algorithms that aims to linearly transform the data such that the resulting dimensions are orthogonal (and therefore can represent the same space), and the dimensions greedily try to explain as much of the total variance as possible (which means the 1st dimension explains the max possible variance any single dimension can, the 2nd explains the most of the remaining variance, and so on). It is a straightforward application of eigenvectors on the matrix $M^TM$.
Calculating the PCA
Why $M^TM$? The mathematical explanation is pretty hefty, but intuitively each value $M^TM_{ij}$ is the dot product of column $i$ with column $j$, which defines a covariance matrix over the columns of $M$. The total variance of this matrix is given by the sum of the diagonal terms in the matrix (commonly called the trace of the matrix).
Orthogonal linear transformations (such as the one performed by the matrix of eigenvectors) cannot increase the total variance. Going back to our aim of maximizing variance as greedily as possible, we want to distribute this sum to lean heavily to the beginning.
A more mathematical formulation of this goal is that PCA aims to solve a series of optimization problems, each one trying to optimize the amount of variance for a single dimension - while still being a linear transformation of the data.
I will not go into the exact math behind it (readers can look to Bishop[1] section 12.1.1 for the proof), but this is equivalent to finding the eigenvalues and the eigenvectors of $M^TM$ and ordering the eigenvectors matrix by their eigenvalues from high to low.
Once this matrix of eigenvectors (let’s call it $T$) is learned, all that’s left is to transform the original matrix $M$, using $P = MT$. As discussed above, this matrix will have the property that its first dimension explains as much of the total variance as possible, the second dimension explains as much of the remaining variance, and so on.
Dimensionality Reduction
The dimensionality reduction comes in the form of dropping some of the eigenvectors from $T$. Notice that $M$ is a matrix of size $n\times m$, $M^TM$ is a matrix of size $m\times m$, and $T$ is a matrix of size $m\times r$, where $r$ is the rank of the matrix $M^TM$. If all rows of $M^TM$ are linearly independent, $r = m$.
If we drop the last few columns of $T$, we will not lose too much variance, as most of the total variance is in the first few columns. Since variance is closely tied to the amount of information in the matrix, dropping the columns with the least variance keeps most of the information. When you take the first $k$ columns of $T$, you get a size $n\times k$ matrix when you apply the transformation, decreasing the dimensionality.
Learned Representation
In terms of Representation Learning, the result of using PCA can be viewed as learning a representation that maximally exposes the underlying variance of the data. This makes separating data into logical pieces (such as classification) easier, as explained in a previous post.
SVD
Singular Value Decomposition is similar to PCA both in terms of its applications and its formulation. It is, however, much more intuitive in terms of Representation Learning, as it explicitly tries to learn the underlying factors (called singular values) in the data, and how the data points and the features relate to those underlying factors.
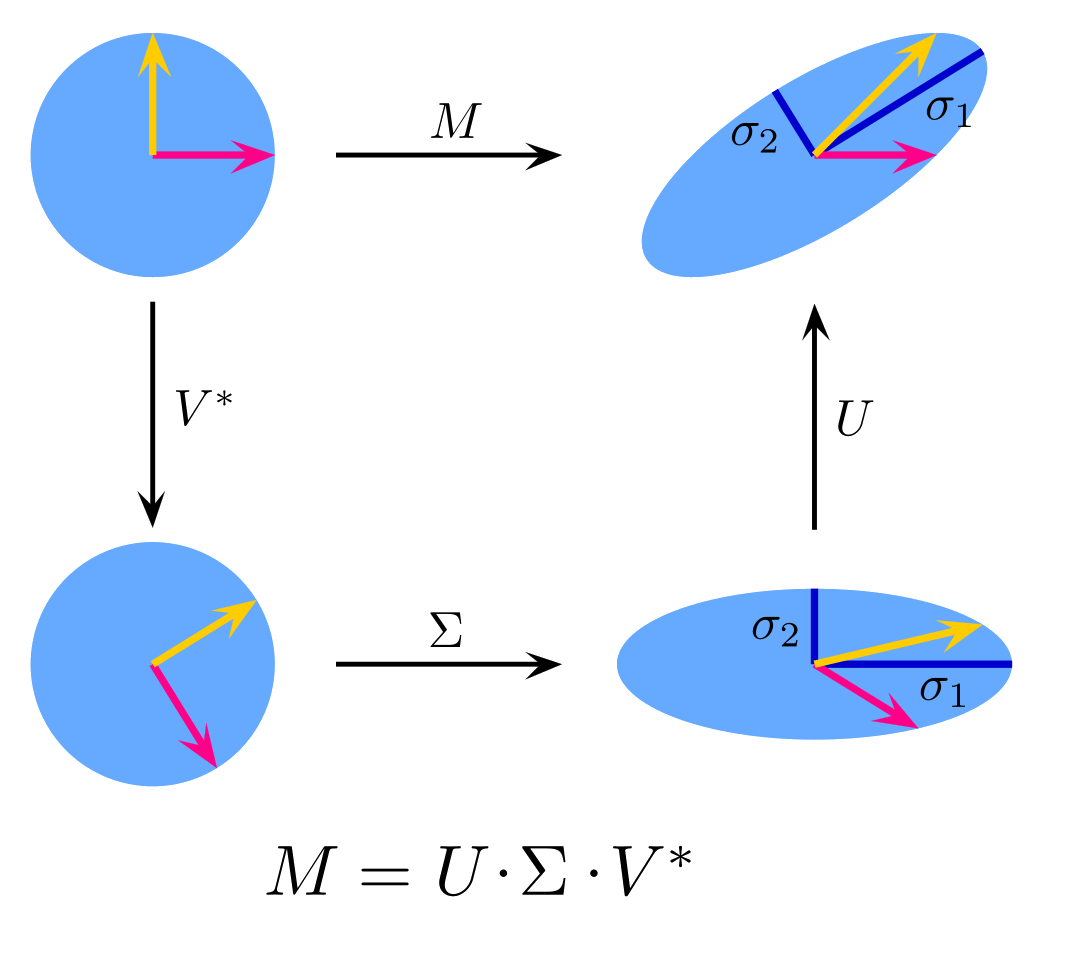
More formally, SVD tries to decompose $M$ into $M = U\Sigma V^T$, where $r$ is the rank of the matrix $M$, $U$ is a matrix of size $n\times r$, $\Sigma$ is a diagonal matrix of size $r\times r$, and $V$ is a matrix of size $m\times r$. Furthermore, $U$ and $V$ are column-orthonormal matrices, such that each of its columns is a unit vector and the dot product of any two columns is 0.
Calculating the SVD
Calculating the SVD requires you to calculate the eigenvectors of $M^TM$ and $MM^T$ again. This is the same as the process explained above, but we can see why that is with a little bit of math:
\[\begin{align} M & = U\Sigma V^T \tag 1 \\ M^T & = V\Sigma^{-1}U^T \tag 2 \\ M^T & = V\Sigma U^T \tag 3 \\ M^TM & = V\Sigma U^TM \tag 4 \\ M^TM & = V\Sigma U^TU\Sigma V^T \tag 5 \\ M^TM & = V\Sigma \Sigma V^T \tag 6 \\ M^TM & = V\Sigma^2V^T \tag 7 \\ M^TMV & = V\Sigma^2V^TV \tag 8 \\ M^TMV & = V\Sigma^2 \tag 9 \end{align}\]Decomposing these equations, equation $(1)$ is the base definition. Equation $(2)$ takes its transpose, which can be simplified into equation $(3)$ because $\Sigma$ is a diagonal matrix. Equation $(4)$ multiplies both sides by $M$ on the right, and equation $(5)$ uses the base definition to extend $M$. Equation $(6)$ uses the fact that $U$ is column-orthonormal, and therefore its multiplication with its transpose gives the identity matrix $I$, which can safely be ignored. Equation $(7)$ simplifies the multiplication of two $\Sigma$ as when you multiply a diagonal matrix by itself, you simply square its values. Finally, equation $(8)$ multiplies both sides by $V$ on the right, and since $V$ is also column-orthonormal, $V^TV$ also gives the identity matrix, which is again ignored to end up with equation $(9)$.
Equation $(9)$ means that $V$ is the matrix of eigenvectors for the matrix $M^TM$. A similar derivation can also be made to obtain $U$ from $MM^T$. Also seen from the same equation, the matrix of eigenvalues for $M^TM$ is the square of the singular value matrix $\Sigma$. We again rearrange the eigenvectors matrix such that $\Sigma$ is sorted in the decreasing order.
Dimensionality Reduction
We can again drop the smallest $k$ eigenvalues, which is equivalent to dropping the last $k$ rows and columns from the $\Sigma$ matrix, and dropping the last $k$ columns from both $U$ and $V$ (note that this drops rows from $V^T$). If all you want is to then use this smaller data, you can use $U\Sigma$ as an input to the rest of your Machine Learning algorithm.
You can also calculate the amount of total variance you’re losing by dropping the last $k$ columns looking at the squares of the singular values. The total variance is the sum of squares of all singular values, and you can calculate which percentage of variance you retain after you drop the last $k$ by summing up the squares of all the remaining singular values and dividing by the original sum.
Learned Representation
As mentioned before, $U$ and $V$ learn to represent the data points and the features in terms of latent factors. If you want to transform a new datapoint $q$ and you want to transform it into a distribution of these latent factors, you only need to compute $qV$, which will give you a vector of size $r$. You can then re-map this vector into the original feature-space using $qVV^T$. This is useful if $q$ has a lot of missing data, which can be estimated through this procedure. This is also the basis of doing recommendations using SVD.
Conclusion
Using SVD in order to expose some of the latent factors is the first step of uncovering the secrets behind your data. Due to the linearity of these operations, SVD cannot learn non-linear representations of data. In the following posts, we will see how we can get rid of this pesky linearity using neural networks.
References: